Using DNA barcodes in the fight against Malaria

Using DNA barcodes in the fight against Malaria
January 27, 2021
By Michelle Lynn D’Souza
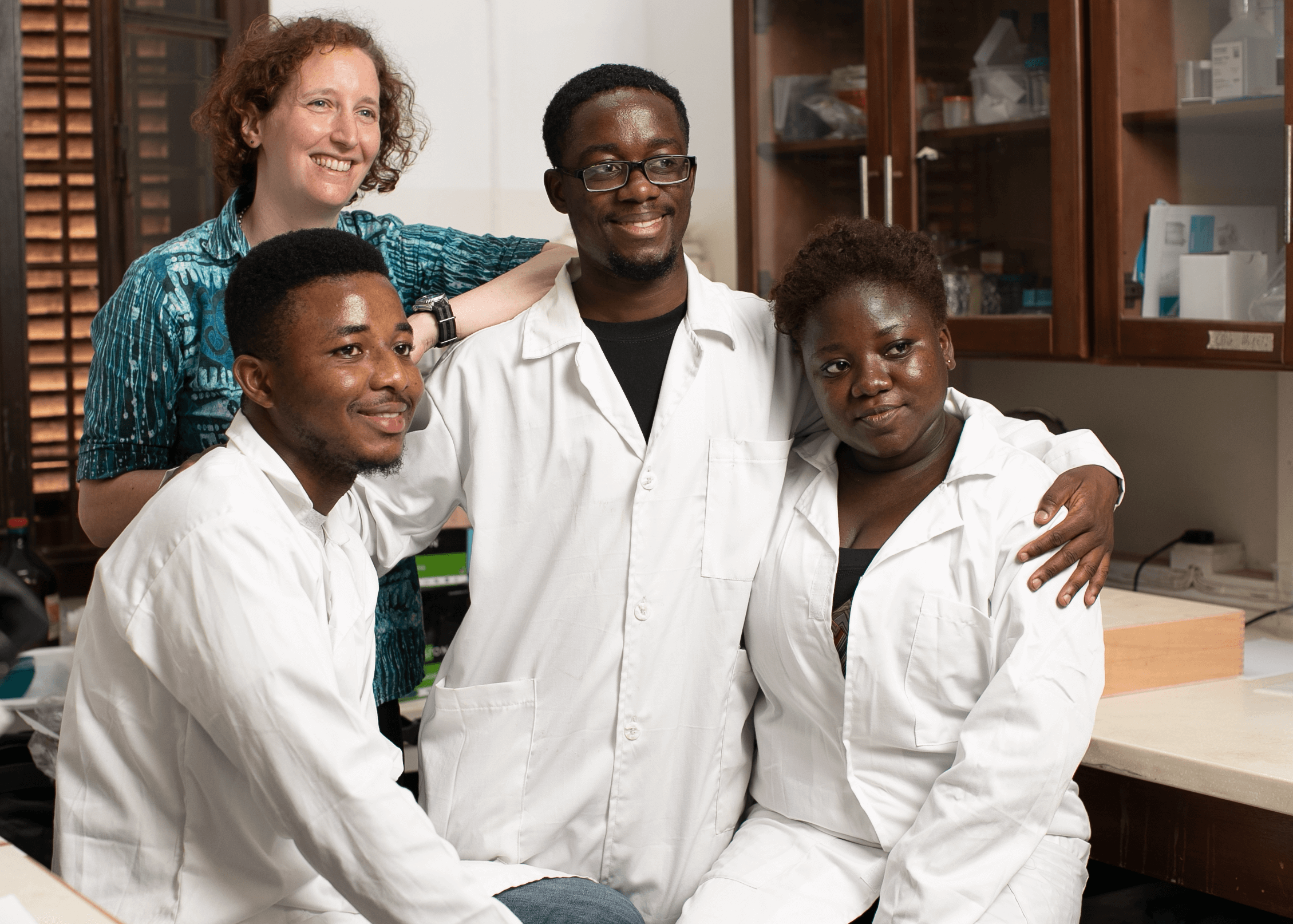
Target Malaria aims to reduce malaria transmission by reducing the population of malaria-transmitting mosquitoes using genetic technologies. As part of their research, they want to know the ecological implications of reducing populations of the mosquito—the key vector for malaria in Africa.
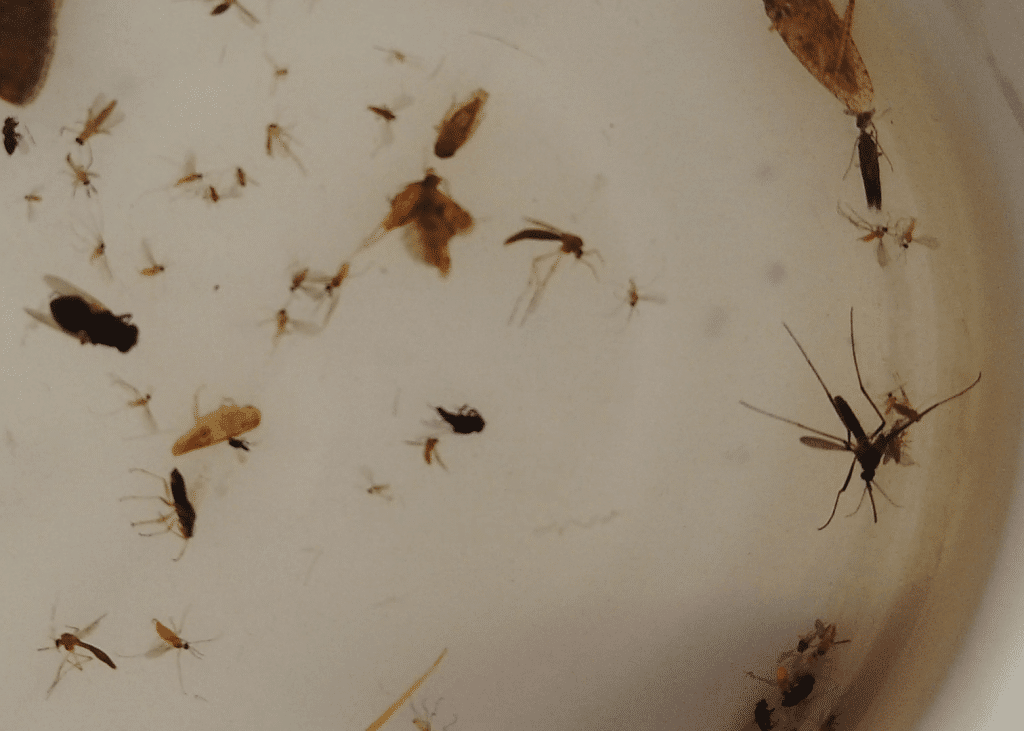
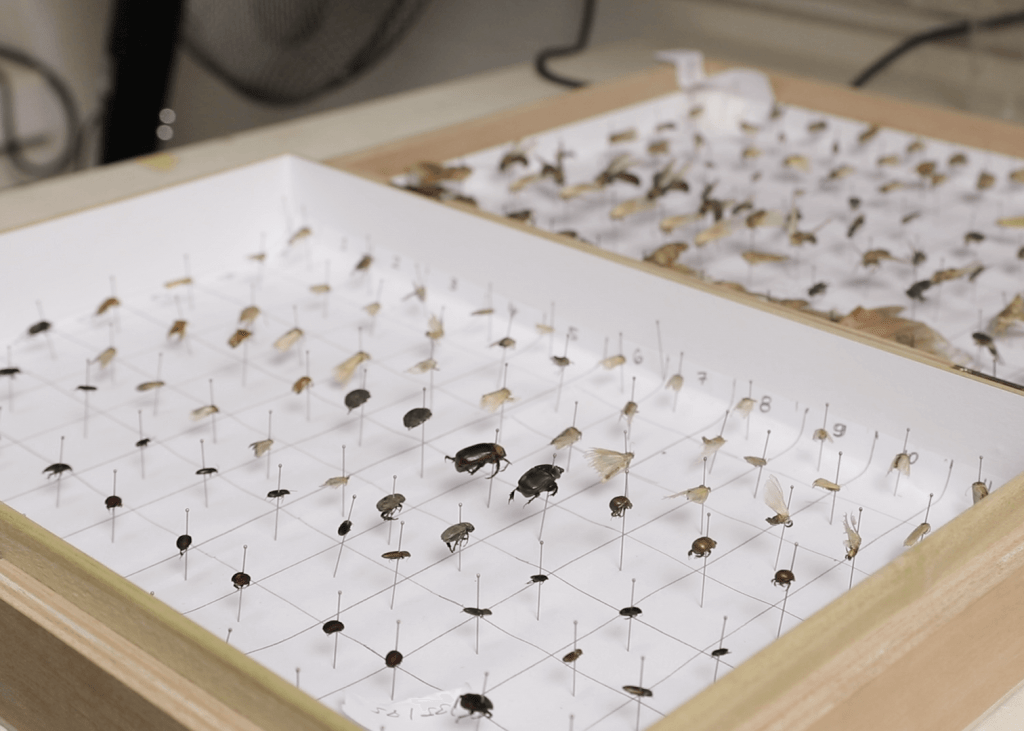
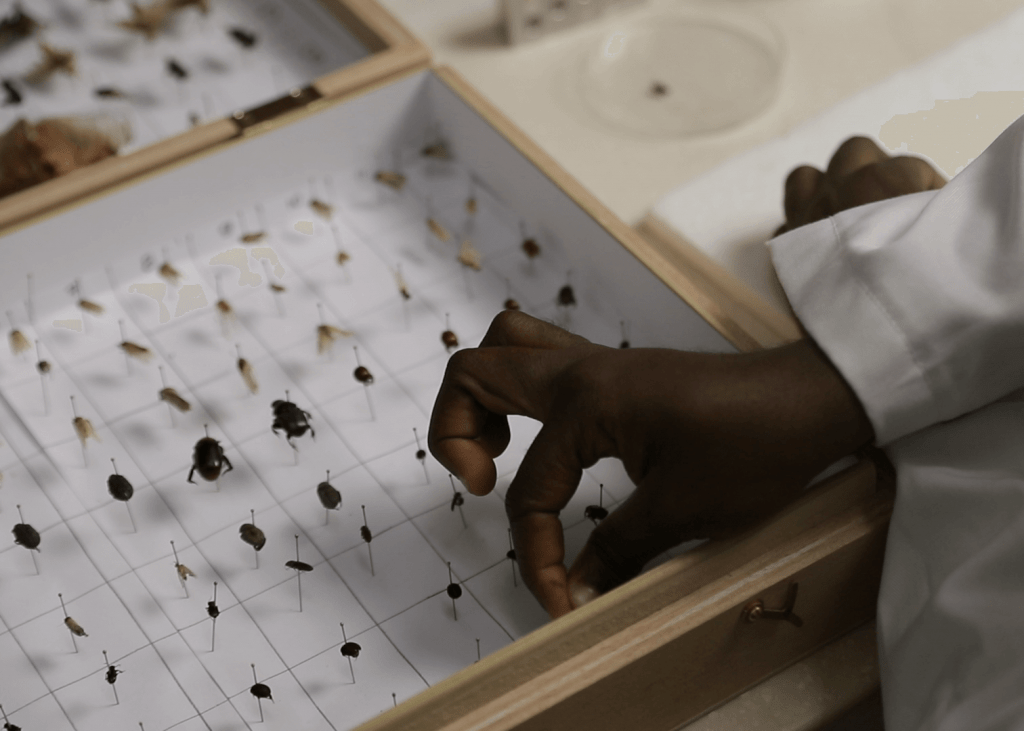
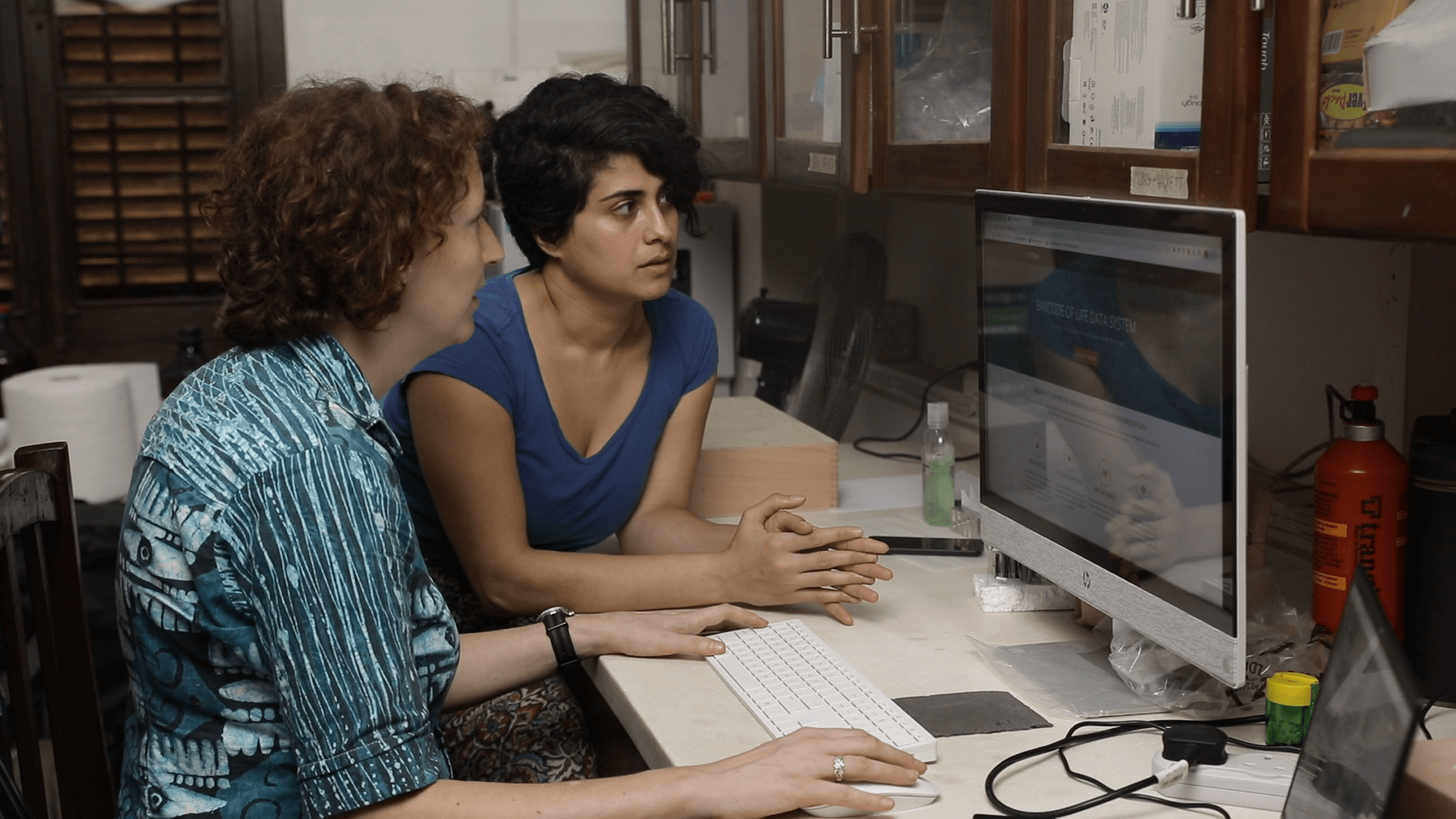
Researchers are using DNA barcoding technologies to catalogue the insect community in Ghana. They will then use this catalogue or library to identify the insect species that are eaten by local birds, bats, and other insect-eating arthropods as well the host species in a mosquito bloodmeal, all through metabarcoding techniques that identify DNA fragments by matching sequences to the local DNA barcode library.
In the end, they will use these data to construct an ecological network that will quantitatively demonstrate how Anopheles gambiae is connected to the other members of the ecosystem.
These efforts are a demonstration of the power of DNA barcoding and its ability to reveal the nature and intensity of interactions among all species. This endeavour, to reveal species interactions to clarify their role in structing biological communities, is a key research theme of BIOSCAN, iBOL’s new seven-year, $180 million global research program that aims to revolutionize our understanding of biodiversity and our capacity to manage it.
For more information on Target malaria efforts in Ghana see: The important interactions behind the itch
For more information on BIOSCAN see: BIOSCAN – Illuminating biodiversity and supporting sustainability
Don't Miss Out!
Subscribe to the iBOL Barcode Bulletin for updates on DNA barcoding efforts, the iBOL Consortium, and more.
Also in BIOSCAN
BIOSCAN: ILLUMINATING BIODIVERSITY AND SUPPORTING SUSTAINABILITY
HOW A TROPICAL COUNTRY CAN DNA BARCODE ITSELF
comment on this article
The Barcode Bulletin moderates comments to promote an informed and courteous conversation. Abusive, profane, self-promotional, or incoherent comments will be rejected.