The important interactions behind the itch
The potential consequences of reducing Anopheles gambiae mosquitoes to control MalariaGhanaian researchers sorting insect samples at the University of Ghana, Accra, Ghana.
IMAGE CREDIT: Michelle L. D’Souza; PHOTO CREDIT: Lema Concepts Africa
Written by

Talya D. Hackett
Department of Zoology, University of Oxford, Oxford

Karen E. Logan
Target Malaria, Dept Life Sciences, Imperial College London, UK

Michelle L. D'Souza
Centre for Biodiversity Genomics, University of Guelph, Guelph, CA
doi:10.21083/ibol.v11i1.6267
Only four months after China reported its first COVID-19 case to the World Health Organization (WHO) the virus had spread to every nation on the African continent. Despite being home to 17% of the world’s population, Africa currently accounts for just 2.5% of COVID-19 related deaths1. But the pandemic may well have caused many more to die, not from coronavirus, but from malaria.
The 2020 World Malaria Report warned that disruptions to malaria prevention and treatment caused by the coronavirus could see as many as 100,000 malaria-related deaths in Sub-Saharan Africa2. Similar effects were unfortunately experienced during the 2014-2015 outbreaks of Ebola in West Africa3. While most strategies being employed to control malaria have worked well, progress to reduce its incidence has stagnated. New strategies are needed to prevent the mortality rate from increasing further and to better prepare countries in the face of other unexpected pressures.
With the ambitious goal to create a world free of malaria, one not-for-profit research consortium–Target Malaria–is developing novel technologies using genetic modification to control the numbers of the malaria-transmitting mosquitoes.
The nature of malaria and the microbe responsible
Malaria is a disease that starts with a small single-celled parasite. This microorganism belongs to the genus Plasmodium, and of the four species that threaten humans, P. falciparum and P. vivax are the most common, and the former the most dangerous4.
Female mosquitoes alone spread malaria in nature. An infected mosquito injects a small number of parasites into its victim’s bloodstream while it feeds, and the parasites then travel to the liver where they multiply rapidly before infecting red blood cells. Flu-like symptoms begin when the parasites break out of the blood cells, one to four weeks after the bite.
Of the 229 million confirmed malaria cases worldwide in 2019, 94% occurred in Africa2. Even more devastating, of the 409,000 malaria-associated deaths, 84% occurred in children under the age of five.
While there are more than 3,500 species of mosquito worldwide and 837 in Africa, three very closely related species are responsible for most transmission of the disease: Anopheles gambiae, Anopheles coluzzii, and Anopheles arabiensis. These three species belong to the Anopheles gambiae complex which, if targeted, is likely to have the largest effect on the transmission of malaria. This species complex has tightly evolved with humans and is the key vector for malaria in sub-Saharan Africa.
Current vector control tools such as insecticides, bed nets, and drugs have been effective in reducing malaria cases but not in eradicating the disease. Target Malaria’s approach is meant to be complementary to the existing interventions by focusing on malaria control by mosquito control.
Researchers unify under Target Malaria
In 2003, Prof. Austin Burt published a seminal paper5 describing the principle of genetically modifying a population of mosquitoes for applications in the control of vector-borne diseases. Prof. Burt predicted that malaria-transmitting mosquitoes could potentially disappear in an area within two years when using these novel genetic tools. It was fortuitous that only a few years prior in Imperial College London, where Prof. Burt was working, a team lead by Prof. Andrea Crisanti had created the first reliable system for germline transformation of a malaria-transmitting mosquito6.
The two groups were brought together in 2005 with a grant given as part of the Grand Challenges in Global Health initiative and in a little over a decade the team’s scientific progress had resulted in a new mechanism for genetic control measures within An. gambiae7.
The initial group of researchers has now grown into a team of 180 project members, with collaborating research partners in Africa, Europe, and North America. As the work progressed, the project grew under its new brand–Target Malaria.
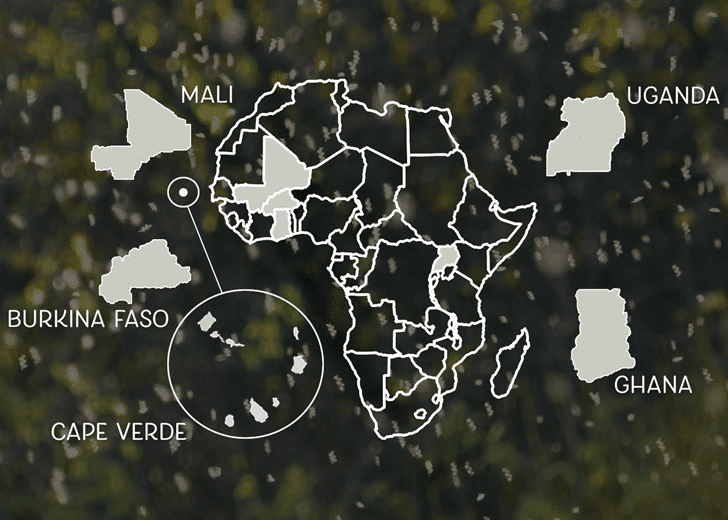
Target Malaria is working with five African partner sites.
IMAGE CREDIT: Michelle L. D’Souza
Today, Target Malaria is working with five African partner sites: Burkina Faso, Cape Verde, Ghana, Mali, and Uganda. This work is being headed by Dr. Abdoulaye Diabate at the Institut de Recherche en Sciences de la Santé (Research Institute on Health Sciences), Bobo-Dioulasso, Burkina Faso; Dr. Adilson de Pina at the Instituto Nacional da Saude (National Institute of Public Health, CCS-SIDA), Cape Verde; Dr. Fred Aboagye-Antwi at the University of Ghana, Accra, Ghana; Dr. Mamadou Coulibaly at the University of Bamako Malaria Research and Training Center, Bamako, Mali; and Dr. Jonathan Kayondo at the Uganda Virus Research Institute, Entebbe, Uganda.
Each of the African partners and their teams bring a different skillset to the collaboration, none being able to deliver all key elements independently. A relationship of co-development is fostered within the project with scientists working together across countries and with communities towards a common objective and vision–a world free of malaria.
The three key guiding pillars
Target Malaria’s work is structured around three key pillars: science, stakeholder engagement, and regulatory affairs. Each pillar is essential for the project’s success, supporting responsible research and development of genetic technologies, a commitment to engaging a wide variety of stakeholders, as well as ensuring compliance with all national regulations and laws.
Your Title Goes Here
Your content goes here. Edit or remove this text inline or in the module Content settings. You can also style every aspect of this content in the module Design settings and even apply custom CSS to this text in the module Advanced settings.
Science
To date, Target Malaria has made significant scientific advancements on the path to developing a new tool for vector control for malaria. Researchers have demonstrated the proof of concept in creation of a transgenic sterile male An. gambiae8 strain, demonstration of the ability to modify a laboratory population of An. gambiae mosquitoes to be male biased9, suppression of a small cage population of laboratory reared An. gambiae mosquitoes10, creation of the first gene drive mosquitoes capable of suppressing a laboratory population of An. gambiae mosquitoes11, modelling the potential of genetic control of malaria mosquitoes12, modelling suppression of malaria vector using gene drive13, and the importation of the first genetically modified mosquitoes (a self-limiting sterile male line) into Burkina Faso for contained laboratory use in 2016 and regulatory approval for the subsequent release of the same self-limiting sterile male line in 2019.
Stakeholder engagement
The list of stakeholders is vast, from grass roots, those local communities where the project is working, through to local civil society organizations, regional governing bodies, and the appropriate governmental agencies, all in-country, as well as a range of interested parties outside of the African partner countries. Target Malaria is committed to ensuring that the stakeholders understand the research and long-term goals of the project enabling them to make an informed decision on whether to support the project’s efforts. Engagement also helps ensure that the research is welcomed and useful in the fight against malaria. Most importantly, Target Malaria will learn a lot from their stakeholders through the process.
Regulatory affairs
While an important aspect of Target Malaria’s strategy is to focus on the communities that might benefit from the technology and that are most concerned by the research activities, they also have an ongoing, transparent dialogue with other stakeholders at the national and international level. For example, the project is taking a phased approach to its development pathway in line with guidance from the WHO14.
Technology being developed
The goal is to develop modified mosquitoes that can pass on to their progeny a self-sustaining genetic change, a process aiming to reduce specific mosquito populations to break the malaria transmission cycle. To do so, Target Malaria is using gene drive, a phenomenon that occurs in nature and causes a selected trait to rapidly increase in frequency through a population via sexual reproduction over several generations. Gene drive works by increasing the likelihood–from the usual 50 per cent to greater than 95 per cent–that a modified gene will be inherited by its offspring. This means that over the course of several generations, a selected trait could become increasingly common within a specific species (depending on the specific area and how the animals move around within it).
Researchers are investigating the use of genes that produce enzymes that cut specific sequences of DNA. Called nucleases, these enzymes found in simple single celled organisms can copy themselves from one chromosome to another5. When introduced into the malaria mosquito, the nuclease works by identifying and cutting a selected site within essential genes targeted by researchers, rendering them functionless, such as reproductive genes. The subsequent effects depend on the nature and importance of the gene.
Target Malaria’s goal is to produce modified malaria mosquitoes that can pass these genes on to greater than 95 per cent of their offspring, so the modification is spread throughout the specific population relatively quickly and is effectively “self-sustaining”. This strategy is known as population suppression, and as the mosquitoes themselves do the work of spreading the modification, it makes the reduction of the malaria mosquito population relatively cost effective and simple to implement.
Current gene drive research is at an early stage, and so definitive decisions about gene drive-based tools are premature. Based on current progress, field releases of a gene drive-based tool are many years away. This gives scientists and stakeholders, specifically those from countries where gene drives might one day be employed, valuable time to consider the important questions of regulation, risk assessment, ethics, and engagement, and to prepare for assessing any application related to gene drive mosquitoes and their potential use as a tool for vector control for malaria.
An ecological approach spearheaded in Ghana
As the gene drive approach in development by Target Malaria will specifically target the An. gambiae complex to reduce its population, it is vital to ensure there are no undesirable consequences to the rest of the plant and animal communities. In Ghana, researchers are focusing on the ecological implications of the work; the role of the An. gambiae mosquito in the broader ecosystem. The ongoing research in Ghana aims to predict these potential effects.
Researchers setting off to sample the insect community around a Ghanaian village.
PHOTO CREDIT: Michelle L. D’Souza
THE ECOLOGICAL ROLE OF MOSQUITO LARVAE IN AQUATIC ENVIRONMENTS
read more…
WHAT WE KNOW, DON’T KNOW, AND THINK WE KNOW ABOUT THE PREDATORS OF MOSQUITOES
read more…
POLLINATORS OR NECTAR THIEVES? THE ROLE OF MALARIA-TRANSMITTING MOSQUITOES IN POLLINATION
read more…
ECOLOGICAL AND EPIDEMIOLOGICAL INSIGHTS FROM BLOOD MEALS
read more…
While some aspects of An. gambiae ecology is well studied, research in Ghana will provide a more complete picture, specifically determining the interactions between An. gambiae and other mosquito species as well as predators, prey, and vertebrate hosts. In this sense, the research is based on a community ecology approach rather than looking at just mosquito ecology.
Researchers are sampling, as far as they are able, insects from the entire aerial communities across all habitats, not just where they expect to find high numbers of An. gambiae. Equally true for insectivores that might feed on mosquitoes or similar small aerial insects, they are taking fecal samples or stomach contents. In aquatic habitats, they are sampling from a range of water bodies and collecting representatives across all insect and insectivorous groups. And, because adult mosquitoes rely on flower nectar for food, they are sampling the pollinator community as well. Finally, they are looking at the community of biting flies and their vertebrate hosts by determining blood-meal interactions to better understand shared hosts and the potential for zoonotic disease transmission15. Importantly, all methods and target sample numbers have been cleared by independent ethics boards at both the University of Ghana and the University of Oxford to ensure there is no lasting impact on the community of plants and animals where the work takes place.
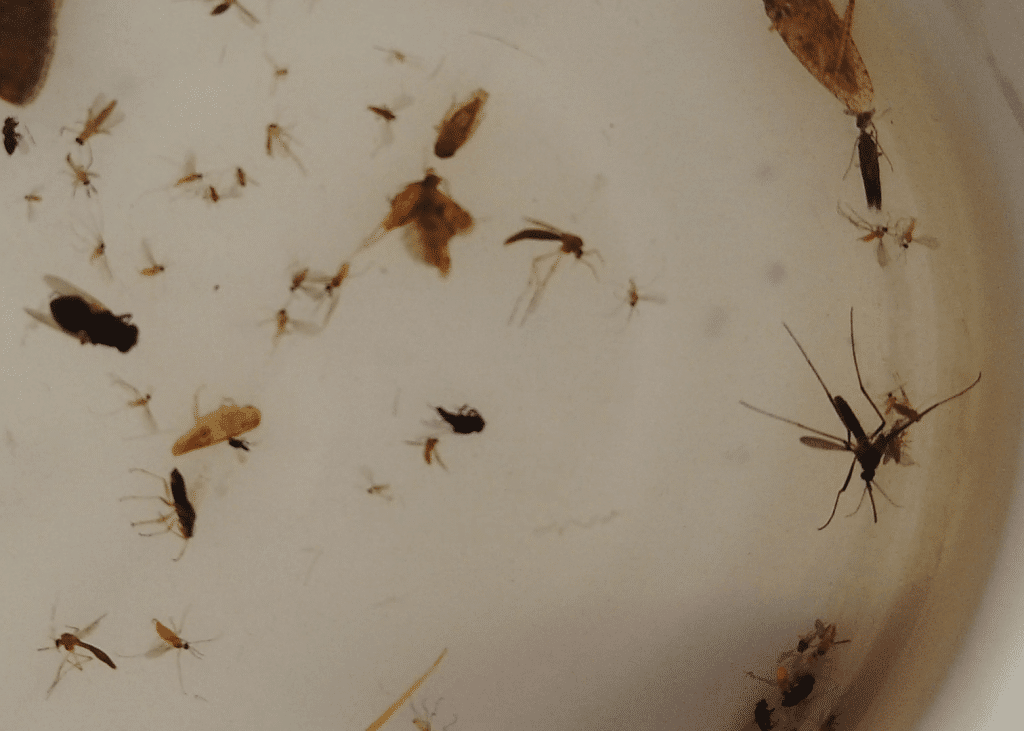
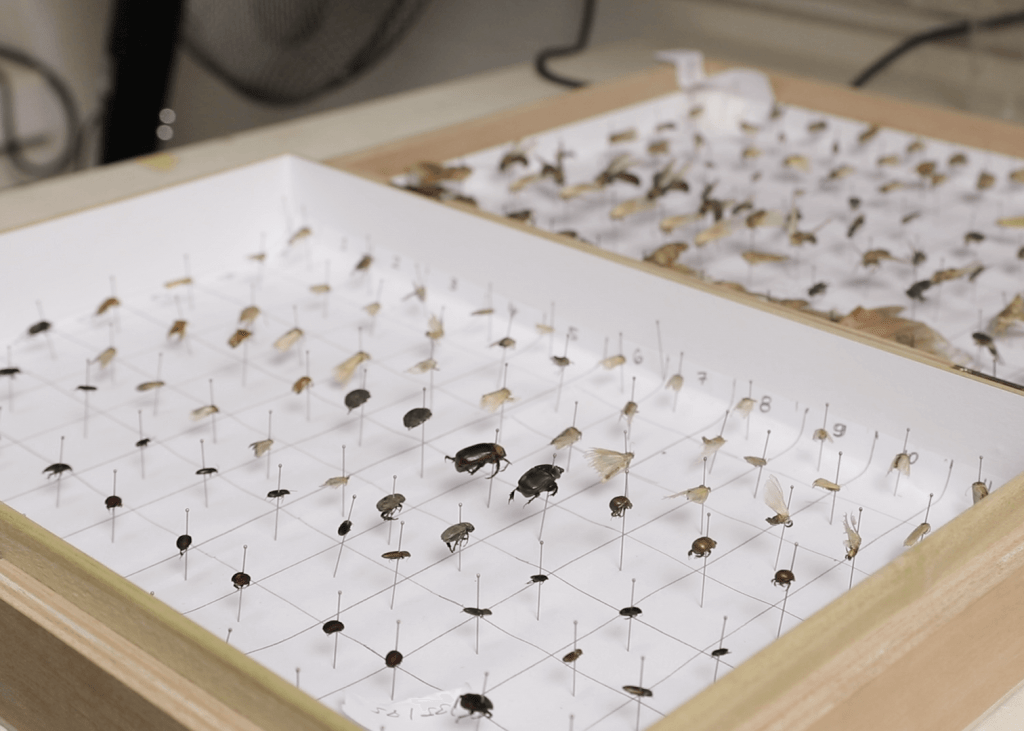
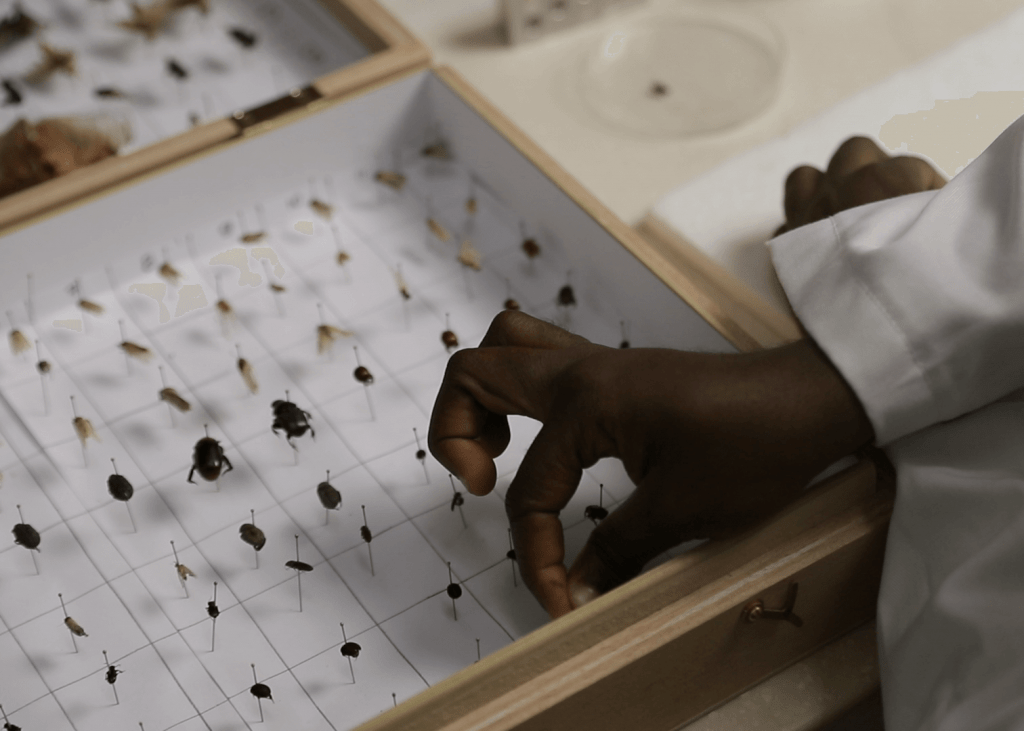



Insect samples collected around a Ghanaian village using Malaise traps (top) before being sorted and pinned (bottom) at the University of Ghana.
PHOTO CREDIT: Lema Concepts Africa
Using this broad community approach allows researchers not only to describe the role of An. gambiae in the ecosystem but also to predict how the rest of the ecological community would respond to An. gambiae reduction. For example, the data would allow them to determine which insects might face more pressure from predators if those that feed on mosquitoes shifted their feeding behaviour to replace An. gambiae in their diet and how this change might affect the rest of the food web. As there is no known animal or plant that relies solely on An. gambiae16, and food webs tend to rewire following minor perturbations17,18, it is predicted that there will not be any significant effects because of An. gambiae population reduction. Regardless, it is necessary to ensure this is the case and a community ecology approach will make this possible.
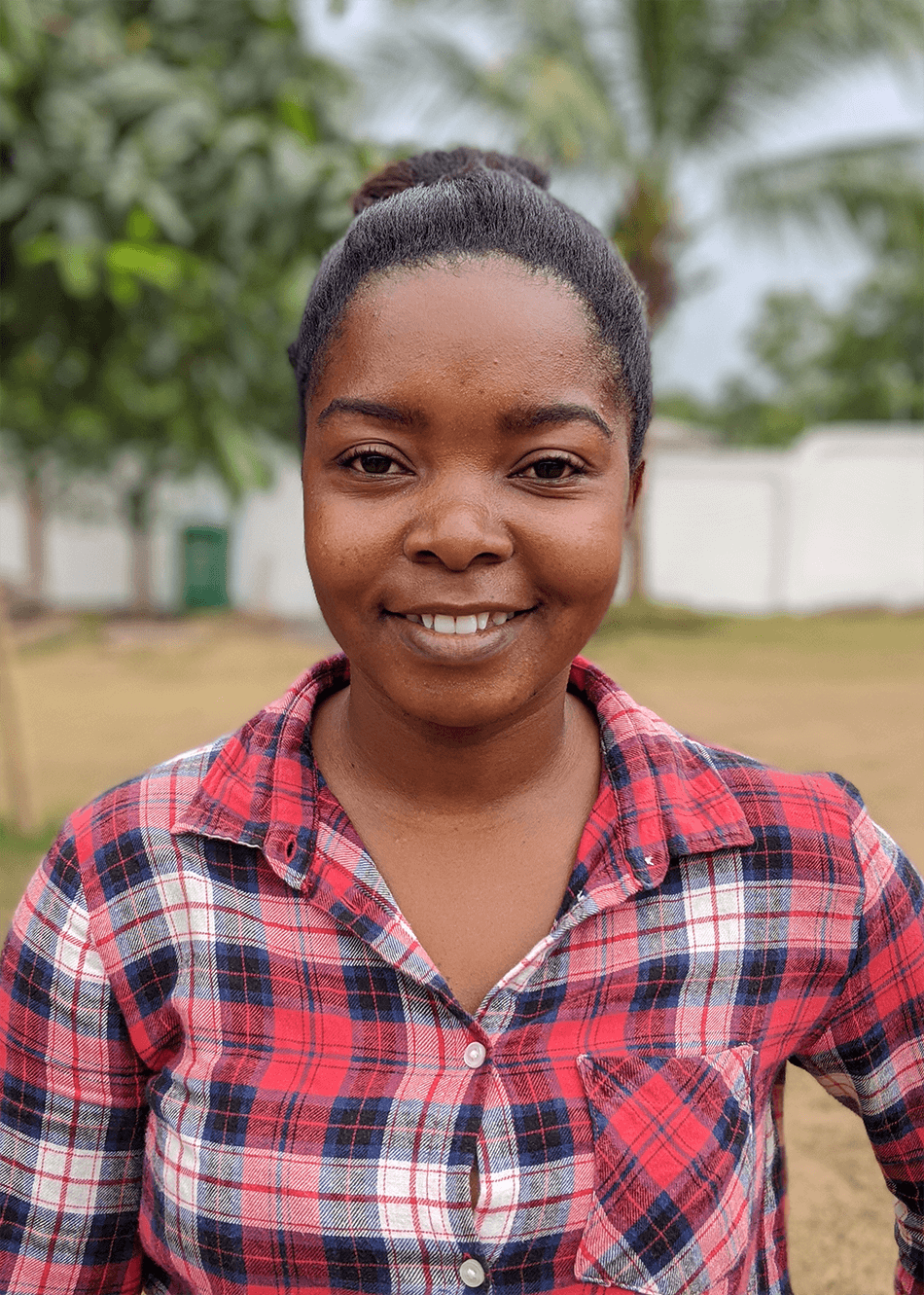
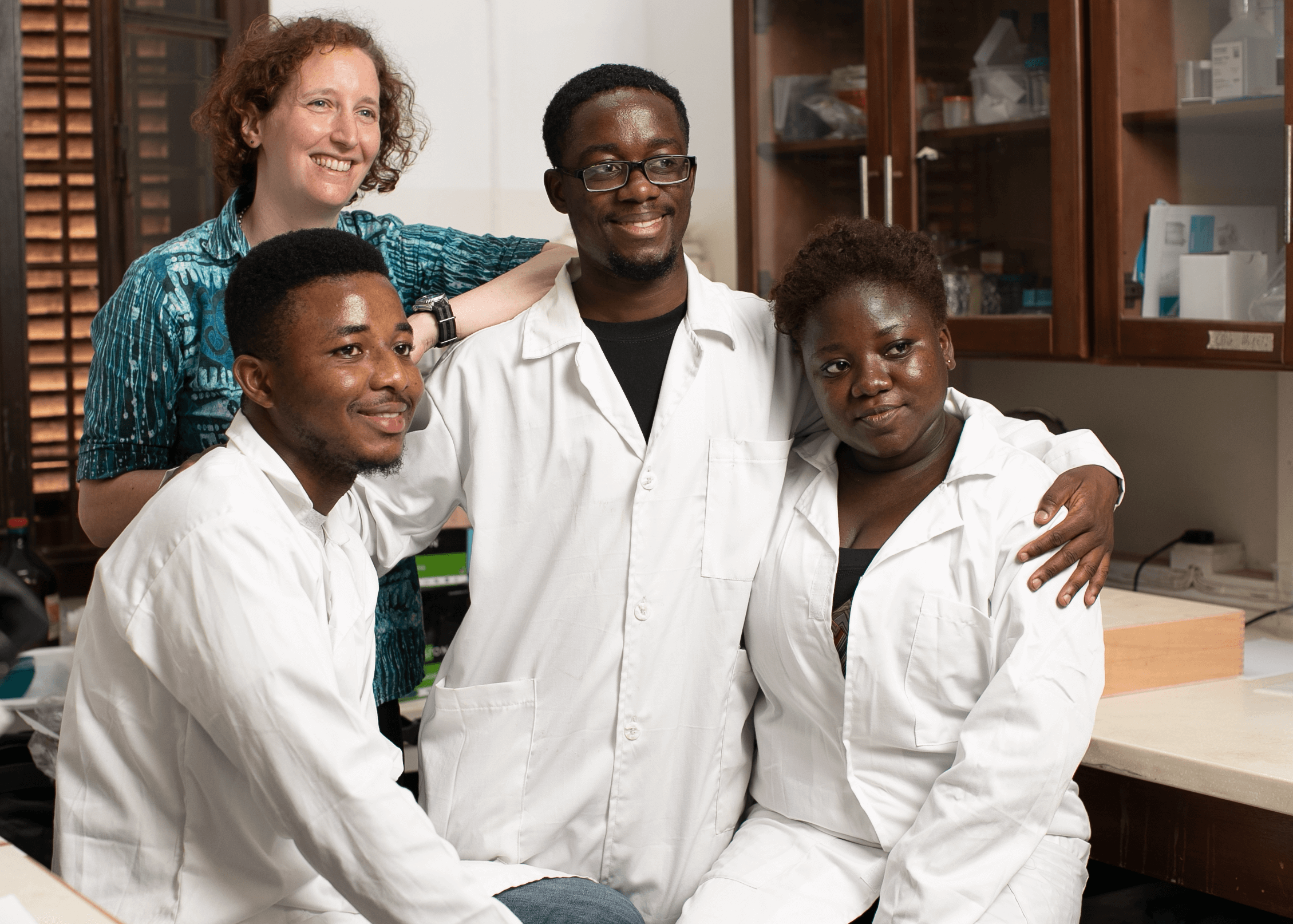




Some of the faces of the research team in Ghana. Dr. Fred Aboagye-Antwi (Ghanaian Principle Investigator, top left), Helen Selorm Wohoyie (Assistant Stakeholder Engagement and Communications Advisor, top center), Divine Dzokoto (Senior Stakeholder Engagement and Communications Advisor, top right) and Dr. Talya D. Hackett (project coordinator), Bernard Aiye Adams, Ezekiel Yaw Donkor, and Naa Na Afua Acquaah (laboratory technicians) (bottom, left to right).
PHOTO CREDIT: Lema Concepts Africa
Building a DNA barcode reference library for Ghana
The tools being used to construct the food web are mostly molecular-based, all requiring the creation of a DNA barcode library for insects in the area as the important first step. To this end, researchers in Ghana regularly collected terrestrial insects from villages in the southeast of Ghana for a year.
Once the library is established, they can then start looking at the feces and the gut contents of insectivores and use that library to match and identify prey DNA fragments using DNA metabarcoding.
Researchers are using a similar metabarcoding approach for the aquatic food web while for the pollination network they will use a combination of traditional observational methods and DNA metabarcoding of pollen from caught insects. Finally, for the blood meal analysis, they are metabarcoding the blood meals of fed mosquitoes and other biting flies to identify what has been bitten.
The ecological research in Ghana is a collaboration between Dr. Fred Aboagye-Antwi at the University of Ghana, Prof. Sir Charles Godfray and Prof. Owen Lewis at the University of Oxford, as well as an extensive team of postdoctoral researchers, Ph.D. students, and technicians at both institutions. Part of the UK team, postdoctoral researcher and project coordinator, Dr. Talya D. Hackett is organizing efforts between countries, including a collaboration with the Centre for Biodiversity Genomics (CBG) in Guelph, Canada, global leader in the field of DNA barcoding. Supported by Prof. Paul D. N. Hebert, the Director of CBG, and Dr. Michelle L. D’Souza, samples from Ghana have begun making their way to the large sequencing platforms housed at the CBG.
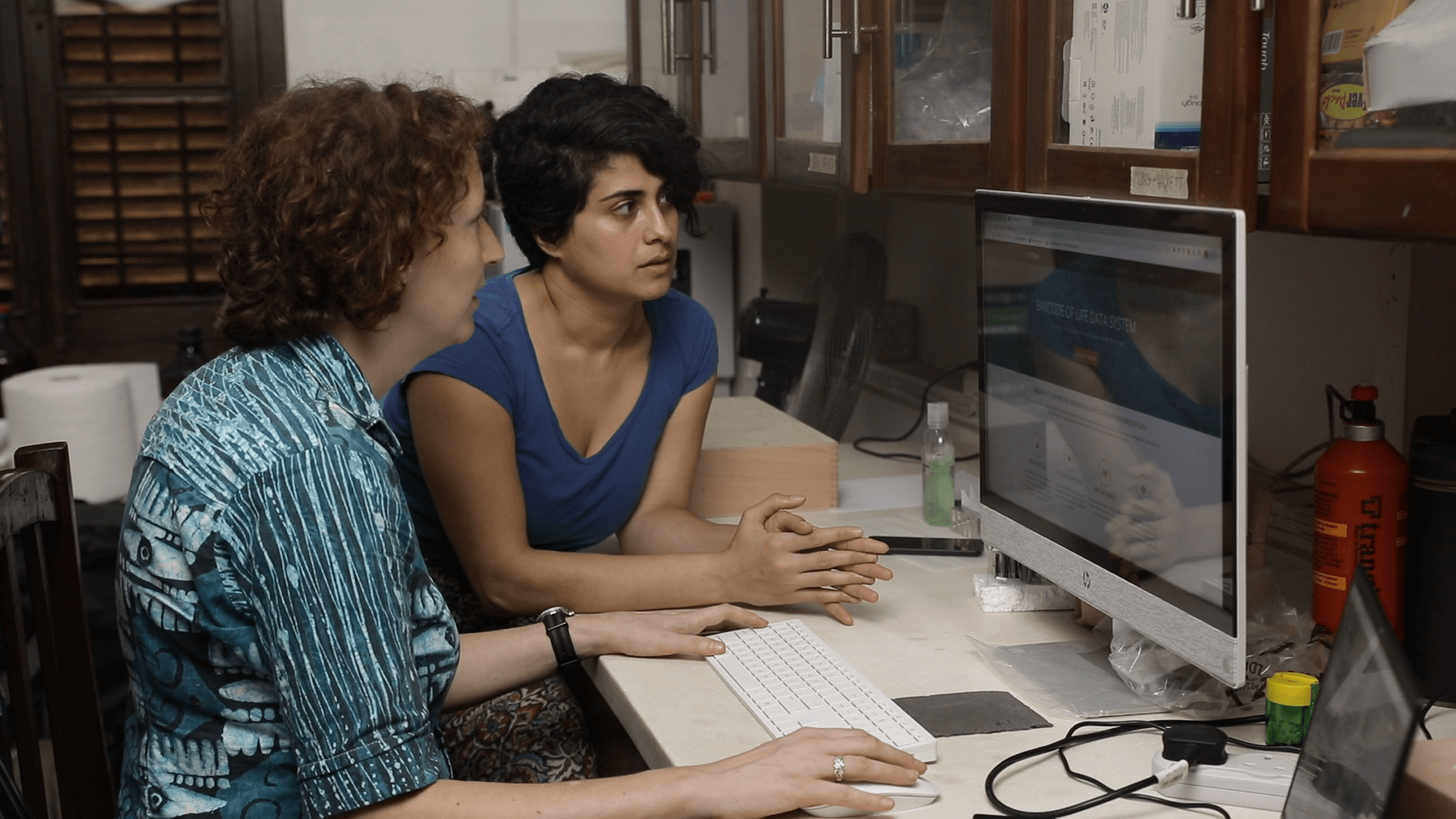

Dr. Talya D. Hackett (left) and Dr. Michelle L. D’Souza (right) discuss DNA barcode data compatibility across platforms EarthCape and BOLD Systems at the University of Ghana.
PHOTO CREDIT: Lema Concepts Africa
So far, about 3,000 insects have been processed and 530 BINs (species proxies) have been documented, about 70% of which are unique to the project. Efforts will ultimately barcode 100,000 specimens and fill a large gap in barcode data currently missing from West Africa.
Conclusions
Examining diets to determine species-specific interactions in a complex community food web is only possible at this large scale with molecular techniques, and only recently, because the costs of DNA barcoding and metabarcoding techniques have dropped. Even five years ago this sort of a project would not have been feasible.
Apart from building large, comprehensive food webs, these data can further inform our understanding of things like community structure and insect abundances across time and space, the dietary overlap of insectivorous species, and niche overlap of different mosquito species.
All data will be made publicly available. Ultimately, this project is creating a wealth of information, not just for Target Malaria’s research goals, but for the broader scientific community and for other people within Ghana and West Africa.
These efforts are a demonstration of the power of DNA barcoding and its ability to reveal the nature and intensity of interactions among all species. This endeavour, to reveal species interactions to clarify their role in structuring biological communities, is a key research theme of BIOSCAN, iBOL’s new seven-year, $180 million global research program that aims to revolutionize our understanding of biodiversity and our capacity to manage it.
If you would like to know more about Target Malaria, go to www.targetmalaria.org
References:
1. World Health Organization (2020) WHO Coronavirus Disease (COVID-19) Dashboard. Accessible at: https://covid19.who.int/
2. World Health Organization (2020) World malaria report. Accessible at: https://www.who.int/teams/global-malaria-programme/reports/world-malaria-report-2020
3. Wang J et al. (2020) Preparedness is essential for malaria-endemic regions during the COVID-19 pandemic. The Lancet 395(10230), 1094–1096. doi: 10.1016/s0140-6736(20)30561-4
4. Crutcher JM, Hoffman SL. Malaria. In: Baron S, editor. Medical Microbiology. 4th edition. Galveston (TX): University of Texas Medical Branch at Galveston; 1996. Chapter 83.
5. Burt A (2003) Site-specific selfish genes as tools for the control and genetic engineering of natural populations. R. Soc. London. Ser. B Biol. Sci. 270:921–928. doi: 10.1098/rspb.2002.2319
6. Catteruccia F, Nolan T, Loukeris TG, Blass C, Savakis C, Kafatos FC, Crisanti A (2000) Stable germline transformation of the malaria mosquito Anopheles stephensi. Nature 405(6789):959-962. doi: 10.1038/35016096.
7. Windbichler N, Menichelli M, Papathanos PA, Thyme SB, Li H, Ulge UY, Hovde BT, Baker D, Monnat RJ Jr, Burt A, Crisanti A. (2011) A synthetic homing endonuclease-based gene drive system in the human malaria mosquito. Nature. 473(7346):212-215. doi: 10.1038/nature09937.
8. Windbichler N, Papathanos PA, Catteruccia F, Ranson H, Burt A, Crisanti A (2007) Homing endonuclease mediated gene targeting in Anopheles gambiae cells and embryos. Nucleic Acids Res. 35(17):5922-5933. doi:10.1093/nar/gkm632
9. Galizi R, Doyle LA, Menichelli M, Bernardini F, Deredec A, Burt A, et al. (2014) A synthetic sex ratio distortion system for the control of the human malaria mosquito. Commun., 5: 1–8.
10. Hammond AM, Kyrou K, Bruttini M, North A, Galizi R, Karlsson X, et al. (2017). The creation and selection of mutations resistant to a gene drive over multiple generations in the malaria mosquito. PLOS Genet. 13:e1007039.
11. Kyrou K, Hammond AM, Galizi R, Kranjc N, Burt A, Beaghton AK, et al. (2018) A CRISPR–Cas9 gene drive targeting doublesex causes complete population suppression in caged Anopheles gambiae mosquitoes. Biotechnol. 36:1062–1071
12. North AR, Burt A & Godfray HCJ (2019) Modelling the potential of genetic control of malaria mosquitoes at national scale. BMC Biol. 17: 26.
13. North AR, Burt A & Godfray HCJ (2020) Modelling the suppression of a malaria vector using a CRISPR-Cas9 gene drive to reduce female fertility. BMC Biol. 18:98.
14. World Health Organization (2014) Guidance framework for testing of genetically modified mosquitoes. Accessible at: https://www.who.int/tdr/publications/year/2014/en/
15. Bellekom B, Hackett TD & Lewis OT (2021) A Network Perspective on the Vectoring of Human Disease. Trends Parasitol.
16. Collins CM, Bonds JAS, Quinlan MM & Mumford JD (2018) Effects of the removal or reduction in density of the malaria mosquito, Anopheles gambiae s.l., on interacting predators and competitors in local ecosystems. Vet. Entomol. 33(1):1–15. doi: 10.1111/mve.12327
17. Timóteo S, Ramos JA, Vaughan IP & Memmott J (2016) High resilience of seed dispersal webs highlighted by the experimental removal of the dominant disperser. Biol. 26:910–915. doi: 0.1016/j.cub.2016.01.046
18. Bartley TJ, McCann KS, Bieg C, Cazelles K, Granados M, Guzzo MM, et al. (2019) Food web rewiring in a changing world. Ecol. Evol. 3: 345–354. doi: 10.1038/s41559-018-0772-3
Don't Miss Out!
Subscribe to the iBOL Barcode Bulletin for updates on DNA barcoding efforts, the iBOL Consortium, and more.
comment on this article
The Barcode Bulletin moderates comments to promote an informed and courteous conversation. Abusive, profane, self-promotional, or incoherent comments will be rejected.


0 Comments