
What can’t be measured won’t be managed: Scientists and U.S. Environmental Protection Agency work together to conserve the Great Lakes
What can’t be measured won’t be managed: Scientists and U.S. Environmental Protection Agency work together to conserve the Great Lakes
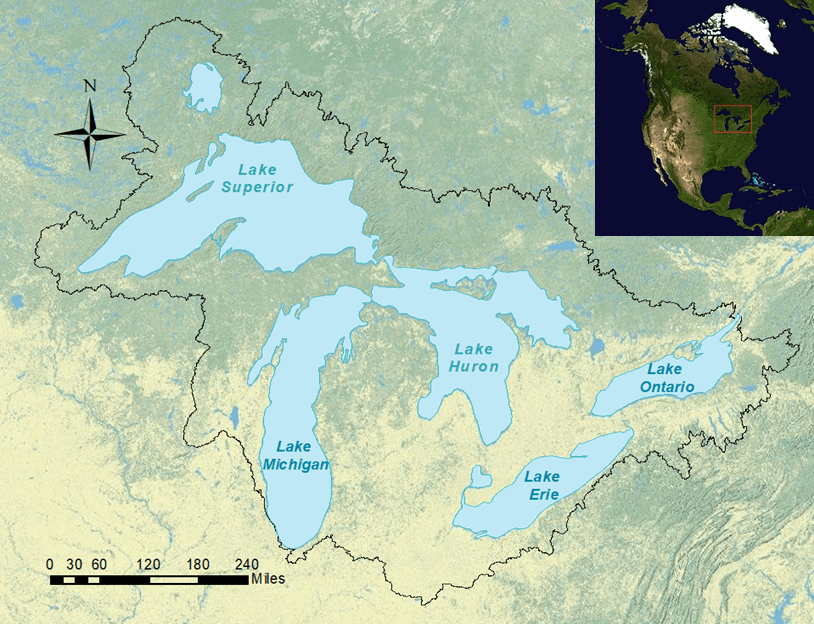
The Laurentian Great Lakes provide extremely valuable ecosystem services to nearly 40 million citizens of Canada and the United States who inhabit the watershed and many other visitors. These lakes are important for commercial navigation and are one of the most valuable freshwater commercial and recreational fisheries in the world. Such heavy use makes them vulnerable to invasive species, of which there are about 180 known to have invaded the five lakes1. However, the lakes’ biodiversity remains startlingly unknown, especially at lower trophic levels, even with strong scientific communities on both sides of the border.
Understanding the impacts of anthropogenic changes on freshwater biodiversity is a major challenge with direct relevance to human health and well-being2. Monitoring and managing aquatic biodiversity needs to involve both academic and government institutions as well as stakeholders spanning farmers, fishermen, global transport companies, and policymakers in order to better inform environmental risk assessment, policy development, and natural resource management. Additionally, evaluating and improving private or public efforts to protect biodiversity requires an ability to quantify biodiversity, beginning with species richness.
The lack of scalable tools for assessing biodiversity has been a major impediment when monitoring the health of freshwater ecosystems. These habitats are dominated by small organisms that are difficult to identify and preserve. Species-level identification based on morphology is often impractical or sometimes even impossible. The process involves expert taxonomists and the special treatment of specimens requires significant investments in money, time, and labour. Therefore, when we rely only on these traditional survey practices, many organisms are identified only to genus/subfamily or simply neglected3,4.
DNA barcoding is a useful tool in these situations because the necessary taxonomic resources can be invested in a more targeted approach once a large number of specimens have been assigned a digital species identifier based on its DNA—the DNA barcode—to create a standardized, reproducible, and scalable solution for monitoring, otherwise difficult to quantify species. By digitizing taxonomic information in the form of a barcode, one needs not taxonomic expertise but simply access to sequencing technology for future identification and monitoring requirements. These technologies are becoming more portable and affordable every day and these tools become even more exciting when we apply non-invasive water sampling to monitor entire fauna from the trace amounts of DNA they leave behind (called ‘environmental DNA’)5.
The Great Lakes Barcoding Project, funded by the United States Environmental Protection Agency (EPA), aims to build a comprehensive genetic barcode library for aquatic invertebrates in the Laurentian Great Lakes watershed. The goal is to improve biodiversity monitoring, provide early detection of non-indigenous species, and inform management efforts to protect biodiversity from threats including climate change, pollution, and invasive species.
At the beginning of the project, only limited genetic information was available for many of the Great Lakes species6. The scale of the Great Lakes and its relatively large invertebrate biodiversity requires this research to be highly collaborative. To this end, the project has brought together several taxonomic experts, molecular ecologists, and aquatic biologists across USA and Canada, from the EPA and research institutions including Cornell University, Buffalo State College, University of Notre Dame, Central Michigan University, and the Centre for Biodiversity Genomics at the University of Guelph.

The Great Lakes DNA Barcoding Project Team: Bret Coggins, Lars Rudstam, Susan Daniel, Adam Frankiewicz, James Watkins, Beth Whitmore, Joe Connolly; bottom row left to right: Sara Westergaard, Michael Pfrender, Bilgenur Baloglu, Kristy Deiner, Ed DeWalt, Alexander Karatayev, Christopher Marshall, Lyubov Burlakova (top to bottom, left to right). In attendance but not pictured: David Lodge, Kara Andres, and Jose Andres. George Rogalskyj and Erik Pilgrim joined electronically.
PHOTO CREDIT: The Great Lakes DNA Barcoding Project
At the end of February 2020, scientists as well as EPA representatives managing or participating in the project gathered at the beautiful Biological Field Station at Cornell University in upstate New York. We shared the latest project updates—everything from taxonomy to biodiversity, from ecological analysis to portable DNA sequencing, and the future of DNA-based monitoring. While the project is still in progress with hundreds of more specimens awaiting analysis, so far, our collaboration has resulted in over 1,000 DNA barcodes spanning over 300 invertebrate species.
This diversity includes more than ten taxonomic classes of invertebrates and is a resource that will improve tracking of non-native and native aquatic species, as well as clarify taxonomic inconsistencies or misrepresentations. The project has stimulated collaborations both within and outside of the main group of researchers and the sharing of specimens, resources, and, most importantly, new ideas and research directions has been an extremely encouraging and productive outcome.
Each plate of specimens sent away for DNA barcode analysis also contains a mix of feelings: satisfaction from a job well done, anticipation of the eventual results, and excitement around the new discoveries that may unfold.
References:
1. Great Lakes Aquatic Nonindigenous Species Information System. Retrieved from: www.glerl.noaa.gov/glansis/index.html
2. IPBES (2019) Global assessment report on biodiversity and ecosystem services of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services. ES Brondizio, J Settele, S Díaz and HT Ngo (editors). IPBES secretariat, Bonn, Germany.
3. Baloğlu B, Clews E and Meier R (2018) NGS barcoding reveals high resistance of a hyperdiverse chironomid (Diptera) swamp fauna against invasion from adjacent freshwater reservoirs. Frontiers in Zoology, 15(1)
4. Srivathsan A, Baloğlu B, Wang W, Tan WX, Bertrand D, Ng AH, Boey EJ, Koh JJ, Nagarajan N and Meier R (2018) A MinION™‐based pipeline for fast and cost‐effective DNA barcoding. Molecular Ecology Resources, 18(5): 1035–1049.
5. Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière‐Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, De Vere N and Pfrender ME (2017) Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Molecular Ecology, 26(21): 5872–5895.
6. Trebitz A, Sykes M, Barge J (2019) A reference inventory for aquatic fauna of the Laurentian Great Lakes. J. Great Lakes Res.
this project is supported by the

Written by

Centre for Biodiversity Genomics, Guelph, ON, Canada

Christopher C. Marshall
Department of Natural Resources, Cornell University, Ithaca, New York, USA
Lars Rudstam
Department of Natural Resources, Cornell University, Ithaca, New York, USA

David M. Lodge
Cornell Atkinson Center for Sustainability and Department of Ecology and Evolutionary Biology, Cornell University, Ithaca, New York, USA
Edward DeWalt
Illinois Natural History Survey, Champaign, Illinois, USA

Paul W. Simonin
Department of Ecology and Evolutionary Biology, Cornell University, Ithaca, New York, USA

Elizabeth Whitmore
Department of Natural Resources, Cornell University, Ithaca, New York, USA
Lyubov Burlakova
Great Lakes Center, Buffalo State College, Buffalo, NY, USA

Kristy Deiner
Department of Environmental Systems Science, ETH Zurich, Zurich, Switzerland
Don't Miss Out!
Subscribe to the iBOL Barcode Bulletin for updates on DNA barcoding efforts, the iBOL Consortium, and more.
comment on this article
The Barcode Bulletin moderates comments to promote an informed and courteous conversation. Abusive, profane, self-promotional, or incoherent comments will be rejected.









